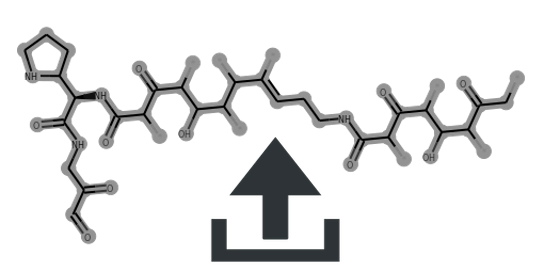
Cluster scaffolds:
1 |
Putative metabolites:
| Name | Rank | Metabolite Score | Similarity | MCS Score | Post-Mod Score | Post-Mod Info | Source | Link | Smiles | Molview | Full Name | Figure | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Xenocyloin A | 0 | 0.57 | 0.53 | 0.75 | 0.0 | Glyco:0/1 * NO2:0/1 | MIBiG | Source | CC(C)C(=O)[C@@H](O)Cc1c[nH]c2ccccc12 | view | Xenocyloin A | ||
| Xenocyloin B | 1 | 0.56 | 0.53 | 0.73 | 0.0 | Glyco:0/1 * NO2:0/1 | MIBiG | Source | CCC(C)C(=O)[C@@H](O)Cc1c[nH]c2ccccc12 | view | Xenocyloin B | ||
| azaspirene | 2 | 0.54 | 0.51 | 0.69 | 0.0 | Glyco:0/1 * NO2:0/1 | MIBiG | Source | CC/C=C/C=C/C1=C(C)C(=O)C2(O1)C(=O)N[C@@](O)(Cc1ccccc1)[C@@H]2O | view | azaspirene | ||
| Xenortide C | 3 | 0.46 | 0.51 | 0.53 | 0.0 | Glyco:0/1 * NO2:0/1 | MIBiG | Source | CN[C@H](C(=O)N(C)[C@@H](Cc1ccccc1)C(=O)NCCc1ccccc1)C(C)C | view | Xenortide C | ||
| Benzylpenicillin | 4 | 0.43 | 0.54 | 0.46 | 0.0 | Glyco:0/1 * NO2:0/1 | MIBiG | Source | CC1(C)S[C@@H]2[C@H](NC(=O)Cc3ccccc3)C(=O)N2[C@H]1C(=O)[O-].[Na+] | view | Benzylpenicillin |